Hepatitis C Virus Dynamics In Vivo: Effect of Ribavirin and Interferon Alfa on Viral Turnover
HEPATOLOGY, July 1998, p. 245-252, Vol. 28, No. 1
Original Articles
Stefan Zeuzem1, Jürgen M. Schmidt2, Jung-hun Lee1, Michael von Wagner1,3, Gerlinde Teuber1, and W. Kurt Roth3
From the 1 Medizinische Klinik II, Klinikum der Johann Wolfgang Goethe-Universität, Frankfurt a.M., 2 Institut für Biophysikalische Chemie der Johann Wolfgang Goethe-Universität, Frankfurt a.M., and 3 Georg-Speyer-Haus, Frankfurt a.M., Germany.
ABSTRACT
Treatment of patients with chronic Hepatitis C with recombinant interferon alfa (rIFN-![]() ) can cause a decrease of serum transaminases and Hepatitis C virus (HCV) RNA. Recent trials evaluating combination therapy of IFN-
) can cause a decrease of serum transaminases and Hepatitis C virus (HCV) RNA. Recent trials evaluating combination therapy of IFN-![]() and ribavirin suggested a potential synergistic effect. From serial measurements of serum HCV RNA concentrations following treatment-induced perturbation of the balance between virus production and clearance, we compared the antiviral efficacy of both IFN-
and ribavirin suggested a potential synergistic effect. From serial measurements of serum HCV RNA concentrations following treatment-induced perturbation of the balance between virus production and clearance, we compared the antiviral efficacy of both IFN-![]() alone and IFN- in combination with ribavirin. Chronically HCV-infected patients were treated with either 3 × 3 MU or 3 × 6 MU rIFN- per week or 3 × 6 MU rIFN-
alone and IFN- in combination with ribavirin. Chronically HCV-infected patients were treated with either 3 × 3 MU or 3 × 6 MU rIFN- per week or 3 × 6 MU rIFN-![]() plus 14 mg/kg of body weight ribavirin per day. The time-dependent HCV RNA concentrations during antiviral treatment were analyzed by iterative least-squares regression. After initiation of antiviral therapy, HCV RNA declined exponentially below the detection limit of the reverse-transcription polymerase chain reaction assay (1,000 HCV RNA molecules per milliliter) in 10 of 26 (39%), 10 of 19 (53%), and 10 of 18 patients (56%) treated with 3 × 3 MU, 3 × 6 MU rIFN-
plus 14 mg/kg of body weight ribavirin per day. The time-dependent HCV RNA concentrations during antiviral treatment were analyzed by iterative least-squares regression. After initiation of antiviral therapy, HCV RNA declined exponentially below the detection limit of the reverse-transcription polymerase chain reaction assay (1,000 HCV RNA molecules per milliliter) in 10 of 26 (39%), 10 of 19 (53%), and 10 of 18 patients (56%) treated with 3 × 3 MU, 3 × 6 MU rIFN-![]() without and with ribavirin, respectively. Viral clearance from serum was faster in patients treated with 3 × 6 MU rIFN-
without and with ribavirin, respectively. Viral clearance from serum was faster in patients treated with 3 × 6 MU rIFN-![]() (t1/2 = 0.23 ± 0.15) compared with patients treated with 3 × 3 MU rIFN-
(t1/2 = 0.23 ± 0.15) compared with patients treated with 3 × 3 MU rIFN-![]() per week (0.67 ± 0.36 days) (P < .004). However, half-lives of viral clearance were similar in patients treated with rIFN-
per week (0.67 ± 0.36 days) (P < .004). However, half-lives of viral clearance were similar in patients treated with rIFN-![]() or rIFN-
or rIFN-![]() plus ribavirin. For virus release from infected hepatocytes, absence and presence of ribavirin yielded half-lives of t1/2 = 2.54 ± 2.10 and t1/2 = 1.99 ± 1.70, respectively, indicating that ribavirin does not significantly inhibit HCV production. In conclusion, the data of the present study indicate that higher rIFN-
plus ribavirin. For virus release from infected hepatocytes, absence and presence of ribavirin yielded half-lives of t1/2 = 2.54 ± 2.10 and t1/2 = 1.99 ± 1.70, respectively, indicating that ribavirin does not significantly inhibit HCV production. In conclusion, the data of the present study indicate that higher rIFN-![]() doses accelerate viral clearance from serum. Ribavirin (14 mg/kg/d), however, lacks synergistic antiviral effects in the treatment of chronic Hepatitis C with 3 × 6 MU rIFN-
doses accelerate viral clearance from serum. Ribavirin (14 mg/kg/d), however, lacks synergistic antiviral effects in the treatment of chronic Hepatitis C with 3 × 6 MU rIFN-![]() per week. (HEPATOLOGY 1998;28:245-252.)
per week. (HEPATOLOGY 1998;28:245-252.)
INTRODUCTION
Treatment of patients with chronic Hepatitis C with recombinant interferon alfa (rIFN-![]() ) can achieve clearance of the Hepatitis C virus (HCV) from serum and liver. However, the overall sustained virological response to IFN-
) can achieve clearance of the Hepatitis C virus (HCV) from serum and liver. However, the overall sustained virological response to IFN-![]() monotherapy is less than 20%.1,2 Additional clinical trials have been conducted to evaluate alternative treatment modalities in chronic Hepatitis C, including therapy with ribavirin. While ribavirin monotherapy revealed no consistent effect on HCV RNA relative to placebo,3,4 results of combination therapy with subcutaneously administered rIFN-
monotherapy is less than 20%.1,2 Additional clinical trials have been conducted to evaluate alternative treatment modalities in chronic Hepatitis C, including therapy with ribavirin. While ribavirin monotherapy revealed no consistent effect on HCV RNA relative to placebo,3,4 results of combination therapy with subcutaneously administered rIFN-![]() and orally administered ribavirin suggested a potential synergistic effect.5-7
and orally administered ribavirin suggested a potential synergistic effect.5-7
Ribavirin (1-![]() -D-ribofuranosyl-1H-1,2,4-triazole-3-carboxamide) is a synthetic purine nucleoside that is structurally similar to guanosine. The drug rapidly enters eukaryotic cells, and after intracellular phosphorylation exhibits virustatic activity against a broad spectrum of DNA and RNA viruses. Several possible mechanisms of action have been proposed, including depletion of the intracellular guanosine triphosphate pools, synthesis of RNA with abnormal 5’cap structures, and inhibition of viral polymerase activity. Furthermore, ribavirin has detectable effects on host immune responses. The detailed mechanism of action, however, is unknown.4,8
-D-ribofuranosyl-1H-1,2,4-triazole-3-carboxamide) is a synthetic purine nucleoside that is structurally similar to guanosine. The drug rapidly enters eukaryotic cells, and after intracellular phosphorylation exhibits virustatic activity against a broad spectrum of DNA and RNA viruses. Several possible mechanisms of action have been proposed, including depletion of the intracellular guanosine triphosphate pools, synthesis of RNA with abnormal 5’cap structures, and inhibition of viral polymerase activity. Furthermore, ribavirin has detectable effects on host immune responses. The detailed mechanism of action, however, is unknown.4,8
In the present study, we treated patients chronically infected with HCV with either 3 × 3 MU rIFN-![]() per week, 3 × 6 MU rIFN-
per week, 3 × 6 MU rIFN-![]() per week, or 3 × 6 MU rIFN-
per week, or 3 × 6 MU rIFN-![]() per week plus 14 mg/kg of body weight ribavirin per day to perturb the balance between virus production and clearance. From serial measurements of changes in viremia in patients responding to antiviral therapy, we obtained kinetic information on the dynamics of HCV replication in vivo. Numerical data modeling was performed to compare the direct antiviral efficacy of the three different treatment schedules.
per week plus 14 mg/kg of body weight ribavirin per day to perturb the balance between virus production and clearance. From serial measurements of changes in viremia in patients responding to antiviral therapy, we obtained kinetic information on the dynamics of HCV replication in vivo. Numerical data modeling was performed to compare the direct antiviral efficacy of the three different treatment schedules.
PATIENTS AND METHODS
In this study, 26 patients chronically infected with HCV were treated with 3 MU rIFN-![]() three times per week subcutaneously. In a different cohort of 37 patients, we administered 6 MU rIFN-
three times per week subcutaneously. In a different cohort of 37 patients, we administered 6 MU rIFN-![]() three times per week subcutaneously. Eighteen patients of this cohort were randomized to receive ribavirin (14 mg per kg of body weight, i.e., 900-1,200 mg) in two divided doses orally per day. Duration of treatment is scheduled for 12 months; the trials are ongoing. All patients were previously untreated, and the diagnosis of chronic Hepatitis C was based on elevated serum transaminase levels, histological examination, and the consistent detection of anti-HCV antibodies (third-generation assay) and HCV RNA. All patients were Hepatitis B surface antigen-negative and negative for the antibody to the human immunodeficiency virus type 1 and type 2. Blood samples were obtained 4 and 1 week before initiation of treatment and subsequently at days 0, 1, 3, 7, 14, 21, 28, and 56. Serum was prepared under a laminar flow bench and frozen at
three times per week subcutaneously. Eighteen patients of this cohort were randomized to receive ribavirin (14 mg per kg of body weight, i.e., 900-1,200 mg) in two divided doses orally per day. Duration of treatment is scheduled for 12 months; the trials are ongoing. All patients were previously untreated, and the diagnosis of chronic Hepatitis C was based on elevated serum transaminase levels, histological examination, and the consistent detection of anti-HCV antibodies (third-generation assay) and HCV RNA. All patients were Hepatitis B surface antigen-negative and negative for the antibody to the human immunodeficiency virus type 1 and type 2. Blood samples were obtained 4 and 1 week before initiation of treatment and subsequently at days 0, 1, 3, 7, 14, 21, 28, and 56. Serum was prepared under a laminar flow bench and frozen at ![]() 80°C. Serum HCV RNA levels were quantified as recently described in detail.9-11 Pretreatment serum HCV RNA levels revealed minor variations (<1 log) in the individual patients, indicating steady-state conditions. Because the viremia level at t = 0 is particularly important for the precision of mathematical modeling, the median pretreatment HCV RNA concentration was applied. Genotyping of HCV (according to the classification of Simmonds et al.12) was performed by reverse hybridization assay (Inno LiPA HCV II, Innogenetics, Ghent, Belgium).13 All patients consented to participate in the study, which was approved by the Ethics Committee for Medical Research in Frankfurt a.M., in accordance with the Declaration of Helsinki. A decline of HCV RNA titer below the detection limit of the reverse-transcription polymerase chain reaction (1,000 copies/mL) within the initial 8 weeks of treatment was observed in 10 of 26 patients treated with 3 × 3 MU rIFN-
80°C. Serum HCV RNA levels were quantified as recently described in detail.9-11 Pretreatment serum HCV RNA levels revealed minor variations (<1 log) in the individual patients, indicating steady-state conditions. Because the viremia level at t = 0 is particularly important for the precision of mathematical modeling, the median pretreatment HCV RNA concentration was applied. Genotyping of HCV (according to the classification of Simmonds et al.12) was performed by reverse hybridization assay (Inno LiPA HCV II, Innogenetics, Ghent, Belgium).13 All patients consented to participate in the study, which was approved by the Ethics Committee for Medical Research in Frankfurt a.M., in accordance with the Declaration of Helsinki. A decline of HCV RNA titer below the detection limit of the reverse-transcription polymerase chain reaction (1,000 copies/mL) within the initial 8 weeks of treatment was observed in 10 of 26 patients treated with 3 × 3 MU rIFN-![]() , 10 of 19 patients treated with 3 × 6 MU rIFN-
, 10 of 19 patients treated with 3 × 6 MU rIFN-![]() , and 10 of 18 patients treated with 3 × 6 MU rIFN-
, and 10 of 18 patients treated with 3 × 6 MU rIFN-![]() and ribavirin (14 mg/kg/d). The remaining patients showed only a transient or no response on serum HCV RNA concentration. Kinetic analyses were only applied to data of treatment responders. Clinical, biochemical, serological, and histological characteristics are summarized in table 1.
and ribavirin (14 mg/kg/d). The remaining patients showed only a transient or no response on serum HCV RNA concentration. Kinetic analyses were only applied to data of treatment responders. Clinical, biochemical, serological, and histological characteristics are summarized in table 1.
| table 1. Demographic, Biochemical, Serological, and Histological Profile of Patients With Chronic Hepatitis C Treated With 3 × 3 MU rIFN- |
An analytical model of the infectious cycle of HCV in vivo has recently been described in which compartment A represents the pool of HCV-infected and noninfected hepatocytes as well as extrahepatic replication sites, BRNA denotes the HCV RNA concentration in serum, and C is a fictitious degradation compartment14:
![]()
HCV is produced in infected hepatocytes and subsequently released into the systemic circulation at rate constant Nk1, where N is the number of virions produced per infected cell, and ln2/k1 is the half-life of virus-producing cells as well as that of virus release. Degradation of free virus from the blood occurs at a rate constant k2. Because antibody-complexed virions may have a rate of degradation different from noncomplexed virions, k2 must be interpreted as a combination of antigen-specific and nonspecific processes. During steady-state conditions before antiviral treatment, adjacent and distant hepatocytes become infected at a rate constant k![]() 1. However, virus uptake by previously uninfected (de novo infection) or infected (superinfection) hepatocytes cannot be discriminated, and transformation of noninfected into virus-producing hepatocytes cannot be measured. After initiation of antiviral therapy, viral clearance is described according to a sequential model given by dBRNA/dt = Nk1 A
1. However, virus uptake by previously uninfected (de novo infection) or infected (superinfection) hepatocytes cannot be discriminated, and transformation of noninfected into virus-producing hepatocytes cannot be measured. After initiation of antiviral therapy, viral clearance is described according to a sequential model given by dBRNA/dt = Nk1 A ![]() (k
(k![]() 1 + k2) BRNA.
1 + k2) BRNA.
As recently described,14 the most significant fit of the observed HCV RNA decline after initiation of therapy is obtained assuming that the predominant antiviral effect of rIFN-![]() is inhibition of de novo infection of susceptible cells (k1 and k2 > 0; k
is inhibition of de novo infection of susceptible cells (k1 and k2 > 0; k![]() 1 = 0). The rate of HCV elimination from serum following initiation of antiviral therapy is then determined by two processes: the clearance of HCV RNA per se and the elimination or suppression of virus-producing cells. Thus, viral RNA data were fitted to a three-compartment sequential-reaction model according to
1 = 0). The rate of HCV elimination from serum following initiation of antiviral therapy is then determined by two processes: the clearance of HCV RNA per se and the elimination or suppression of virus-producing cells. Thus, viral RNA data were fitted to a three-compartment sequential-reaction model according to
![]() (1)
(1)
where k1 and k2 denote the rate constants associated with increase and decrease of viral RNA concentration B, respectively. Accordingly, the differential change in the virus concentration B shall obey the hypothesis
Integration requires an assumption on the time dependence of A be made as well as an initial condition for the compartmental populations be given. Let
indicating the concentration of virus-producing cells leveling off at rate dA/dt = ![]() k1A after the initiation of treatment. Unless supported by dedicated experiments, e.g., cell counts, e absolute concentration of A remains unknown but can be related to the observed quantity B using the ratio N as defined above, thus linking virus-level growth to cell damage. It is also implied that infected and noninfected cells in compartment A cannot be distinguished by exclusively sampling compartment B.
k1A after the initiation of treatment. Unless supported by dedicated experiments, e.g., cell counts, e absolute concentration of A remains unknown but can be related to the observed quantity B using the ratio N as defined above, thus linking virus-level growth to cell damage. It is also implied that infected and noninfected cells in compartment A cannot be distinguished by exclusively sampling compartment B.
The first-order differential equation as recast from the two constraining conditions, Eq. 2 and Eq. 3, is given by
Solution of Eq. 4 involves an auxiliary integrating factor exp(k2t) to be inserted, leading to
where the left-hand side of Eq. 5 is identified as the total derivative
Integration of Eq. 6 within the boundaries 0 and t leads to
which, after elimination of the auxiliary term, recasts into
![]()
From the initial condition dB/dt = 0, the stationary levels before drug administration follow from Eq. 2 as A0 = (k2/Nk 1) B0. Substitution into Eq. 8 finally leads to the equation given by Wei et al. as applied in the modeling of human immunodeficiency virus kinetics15 (for convenience, the plasma viral concentration V and the rate constants a and u in the original literature are identified here with B, k1, and k2, respectively),
This model is valid for any number of virions produced per infected cell, because the factor N has been eliminated. Thus, without experimental evidence, the ratio N as defined (Eq. 2) is finally fully incorporated into either the rate constant k1 or the cell concentration A0.
The model given by Eq. 9 takes two separate processes of elimination of virus-producing cells and free virus associated with k1 and k2, respectively, into account. However, the numerical symmetry in the rate equation precludes the two biological processes to be distinguished. Furthermore, it is assumed that the same rate constant k1 is identical for both release of virions produced by infected cells and loss of virus-producing cells. We emphasize that this compound decay curve from Eq. 9 is unable to fit our observed HCV RNA quantification data of the present study, which are characterized by an initially rapid decline followed by a slower one.
Therefore, we considered a definition of the anyway undetermined compartment A, which avoids the duplicate use of the constraining condition given in Eq. 2 and which is less stringent than that applied for the transformation of Eq. 8 to Eq. 9. A general type of the time dependence of the viral compartment that does not make preliminary assumptions on N is
For N = 1, the model of Wei et al. results. In our model,14
the multipliers to the exponential terms are due to an initial condition in which the number of virions produced per hepatocyte was chosen such that B(0) = A(0) including a rate-dependent ratio N = k2/k1 as rationalized in the following.
The observed decline of serum HCV RNA after initiation of antiviral therapy is dominated by two exponential terms related to influx and outflux at rate constants k1 and k2, respectively. Given that the rates differ, a semilogarithmic representation of the concentration B(t) versus t reveals two tangents intersecting at an angle depending on the relative values of the rate constants. Irrespective of a convex or concave shape of the smoothened log curve in a certain range of transitional t values, the steeper tangent is always associated with the faster process, while the slower process, associated with the smaller rate constant and with the shallower slope, determines the turnover on the long term and is considered rate-limiting. Note that also in our model, rate constant k1 describes both release of virions produced by infected cells and loss of virus-producing cells. The observed HCV RNA concentrations following interferon administration imply(k2 > k1). This is compatible with the biological point of view that an infected hepatocyte must produce more than one virion per lifecycle (N > 1). In addition, a rate-dependent ratio N allows to account for possible differences in HCV release from hepatocytes in infected individuals.
Furthermore, detailed data regression was performed with the proportionality k2/Nk1 adjusted to several orders of magnitude between 0.001 and 1,000. Fit significance was highest for a value of about 1. Median viremia in patients with chronic Hepatitis C is about 107 copies per milliliter.11 Multiplication with the extracellular fluid volume, which is estimated to be 20% of the individual patient’s body weight, and assuming that serum and extracellular fluid compartments are in equilibrium, results in an estimate of B0 of around 1.4 × 1011 copies. The liver contains approximately 2 × 1011 hepatocytes (A0).16 Thus, the highest fit significance for a value of 1 of the proportionality k2/Nk1 appears biologically well supported. Note that this does not imply a production rate of one virion per hepatocyte, because compartment A is defined as the pool of HCV-infected and uninfected hepatocytes and only 5% to 40% of hepatocytes are infected with HCV in the chronic state of the disease.17,18
From experimental values of BtRNA (time-dependent HCV RNA serum concentrations), kinetic parameters B0RNA, k1, and k2 were obtained by iterative least-squares fitting.14 The protocols included a repeatedly initialized down-hill simplex optimization.19 Typically, less than 500 function evaluations were needed for convergence. The agreement between simulated and observed data was characterized in a least-squares sense by the normalized fit-error s2 = ![]() in [(Bi sim
in [(Bi sim ![]() Biobs)/
Biobs)/![]() i]2, where i runs over all n data points and
i]2, where i runs over all n data points and ![]() i is the uncertainty of a sample set to either 5% or to a minimum of 1 × 103 molecules per milliliter in HCV RNA determinations. To account for the intrinsic nonlinear properties of the model function, confidence boundaries for the parameters B0, k1, and k2 were derived from the fractional increase in the sum of squares of residuals20,21 computed according to (smax/smin)2 = 1 + ( p
i is the uncertainty of a sample set to either 5% or to a minimum of 1 × 103 molecules per milliliter in HCV RNA determinations. To account for the intrinsic nonlinear properties of the model function, confidence boundaries for the parameters B0, k1, and k2 were derived from the fractional increase in the sum of squares of residuals20,21 computed according to (smax/smin)2 = 1 + ( p ![]() 1)(n
1)(n ![]() p)
p)![]() 1 F(p
1 F(p ![]() 1, n
1, n ![]() p,
p,![]() ),
), ![]() i where n and p are the number of HCV RNA samples and fit parameters, respectively, and F is the Fisher variance ratio. For the varying numbers of degrees of freedom, critical F values for a two-tailed test of the F distribution function were obtained from statistical tables.22 The rejection probability was set to = (31.7/2)% associated with one standard deviation based on the assumption of a multivariate Gaussian distribution. The deviations in each of the optimum parameters required to increase the sum of squares of the residuals from smin2 to the threshold smax2 were iteratively determined using a modified secant algorithm.23 The asymmetry of the s2 isocontour on the error hyper-surface was tested with a bidirectional search using positive and negative deviations in each single parameter.20
i where n and p are the number of HCV RNA samples and fit parameters, respectively, and F is the Fisher variance ratio. For the varying numbers of degrees of freedom, critical F values for a two-tailed test of the F distribution function were obtained from statistical tables.22 The rejection probability was set to = (31.7/2)% associated with one standard deviation based on the assumption of a multivariate Gaussian distribution. The deviations in each of the optimum parameters required to increase the sum of squares of the residuals from smin2 to the threshold smax2 were iteratively determined using a modified secant algorithm.23 The asymmetry of the s2 isocontour on the error hyper-surface was tested with a bidirectional search using positive and negative deviations in each single parameter.20
Rate constants k3 and k4 denote the release of alanine transaminase (ALT) and aspartate transaminase (AST) from hepatocytes and the degradation of transaminases, respectively. B0ALT, and B0AST are the initial serum transaminase levels before rIFN-![]() administration:
administration:
![]()
Rate constants k3 and k4 were calculated from the time-dependent concentrations in analogy to the HCV RNA kinetics with the difference that the sensitivity of transaminase measurements was set to 1 U/L. Because of membrane permeability and normal regeneration of hepatocytes, a baseline level of serum transaminase activity (B![]() ALT,AST) is observed. Thus, fitting of the experimental data required four parameters in an extended equation according to
ALT,AST) is observed. Thus, fitting of the experimental data required four parameters in an extended equation according to
![]()
Double-exponential models allow in-flux and out-flux of the serum HCV RNA compartment to be separated, in contrast to simple first-order kinetic models,15,24 in which such a discrimination of virus production and clearance is not possible. The time-dependent variation of the viral load after drug administration is given by the first derivative of the integrated rate law for the viral compartment with respect to time,
From the initial transitory relaxation process, viral turnover during the stationary phase before perturbation can be estimated, and in fact has been exploited to elucidate the enormous throughput f viral particles in the chronic phase of infection.15,24 Solved for time t = 0, the initial rate in the model function of Eq. 11 amounts to
while that of Eq. 9 is identically zero. To estimate the minimum value of viral production, we multiplied B0 in a previous study by the respective rate k1,14 which is the more conservative approach because of its rate-limiting effect. Because the rate k1 possesses some duality (scaled virus release from infected cells as well as decline of infected cells) and because virus clearance from serum with rate k2 dominates the initial decline in compartment B after perturbation of the steady state, we decided to use rate k2 for calculating the minimum daily turnover of HCV RNA molecules. Thus, minimum daily virus production and clearance was calculated according to B0RNA k2 multiplied by the extracellular fluid volume, which was estimated to be 20% of the individual patient’s body weight, and assuming that serum and extracellular fluid compartments are in equilibrium.
Typically, the fit significances exceeded 95% as tested by ANOVA. Data are expressed as median or mean ± SD. The ![]() 2 test with Yates’ correction was used for statistical analysis of comparison between group frequencies. The two-sample Student’s t test was applied when continuous variables were considered normally distributed by the Kolmogorov-Smirnov test. P < .05 was considered statistically significant. The analyses were performed using the BiAS statistical software (Department of Biostatistics, University of Frankfurt a.M.).
2 test with Yates’ correction was used for statistical analysis of comparison between group frequencies. The two-sample Student’s t test was applied when continuous variables were considered normally distributed by the Kolmogorov-Smirnov test. P < .05 was considered statistically significant. The analyses were performed using the BiAS statistical software (Department of Biostatistics, University of Frankfurt a.M.).
RESULTS
Patients in the three treatment groups of the present study (3 × 3 MU rIFN-![]() per week, 3 × 6 MU rIFN-
per week, 3 × 6 MU rIFN-![]() per week, 3 × 6 MU rIFN-
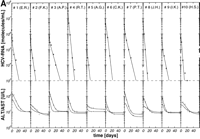
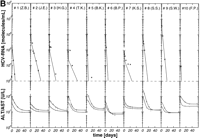
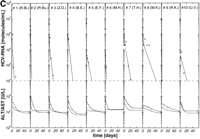
per week, 3 × 6 MU rIFN-![]() per week plus 14 mg/kg ribavirin per day) were well matched regarding age, HCV genotype, pretreatment viremia, and liver histology (table 1). After initiation of antiviral therapy, HCV RNA declined exponentially below the detection limit of the reverse-transcription polymerase chain reaction assay (1,000 HCV RNA molecules per milliliter) in 10 of 26 (39%), 10 of 19 (53%), and 10 of 18 patients (56%) treated with 3 × 3 MU, 3 × 6 MU rIFN-
per week plus 14 mg/kg ribavirin per day) were well matched regarding age, HCV genotype, pretreatment viremia, and liver histology (table 1). After initiation of antiviral therapy, HCV RNA declined exponentially below the detection limit of the reverse-transcription polymerase chain reaction assay (1,000 HCV RNA molecules per milliliter) in 10 of 26 (39%), 10 of 19 (53%), and 10 of 18 patients (56%) treated with 3 × 3 MU, 3 × 6 MU rIFN-![]() without and with ribavirin, respectively (Fig. 1A-1C). Thus, the portion of initial responders was similar in the two treatment groups with 3 × 6 MU rIFN-
without and with ribavirin, respectively (Fig. 1A-1C). Thus, the portion of initial responders was similar in the two treatment groups with 3 × 6 MU rIFN-![]() , but lower in patients treated with 3 × 3 MU rIFN-
, but lower in patients treated with 3 × 3 MU rIFN-![]() per week (P = not significant) (table 1). From the kinetic parameters, the minimum daily production and clearance of HCV was calculated for each case. The median daily turnover was 1.8 × 1011 virions per day (range, 4.5 × 109 to 1.6 × 1013 virions/d).
per week (P = not significant) (table 1). From the kinetic parameters, the minimum daily production and clearance of HCV was calculated for each case. The median daily turnover was 1.8 × 1011 virions per day (range, 4.5 × 109 to 1.6 × 1013 virions/d).
As recently described in patients treated with 3 × 3 MU rIFN-![]() subcutaneously per week14 and confirmed in the present study for patients treated with either 3 × 3 MU or 3 × 6 MU rIFN-
subcutaneously per week14 and confirmed in the present study for patients treated with either 3 × 3 MU or 3 × 6 MU rIFN-![]() per week, the most significant fit of the observed HCV RNA decline after initiation of therapy is obtained assuming that the predominant antiviral effect of rIFN-
per week, the most significant fit of the observed HCV RNA decline after initiation of therapy is obtained assuming that the predominant antiviral effect of rIFN-![]() is inhibition of de novo infection of susceptible cells (k1 and k2 > 0; k
is inhibition of de novo infection of susceptible cells (k1 and k2 > 0; k![]() 1 = 0). Results for virus release from infected cells in patients treated with 3 × 3 MU or 3 × 6 MU rIFN-
1 = 0). Results for virus release from infected cells in patients treated with 3 × 3 MU or 3 × 6 MU rIFN-![]() per week yielded similar half-lives of t1/2 = ln2/k1 = 2.28 ± 1.17 and 2.54 ± 2.10 days, respectively (table 2). The half-lives of viral clearance from serum were significantly shorter in patients treated with 3 × 6 MU rIFN-
per week yielded similar half-lives of t1/2 = ln2/k1 = 2.28 ± 1.17 and 2.54 ± 2.10 days, respectively (table 2). The half-lives of viral clearance from serum were significantly shorter in patients treated with 3 × 6 MU rIFN-![]() compared with patients treated with 3 × 3 MU rIFN-
compared with patients treated with 3 × 3 MU rIFN-![]() per week (t1/2= ln2/k2 = 0.23 ± 0.15 vs. 0.67 ± 0.36 days; P < .004) (table 2), suggesting a dependence of the rate constant k2 on the IFN-
per week (t1/2= ln2/k2 = 0.23 ± 0.15 vs. 0.67 ± 0.36 days; P < .004) (table 2), suggesting a dependence of the rate constant k2 on the IFN-![]() dose, while rate constant k1 remains essentially unchanged.
dose, while rate constant k1 remains essentially unchanged.
| table 2. Kinetic Data of HCV and ALT and AST Turnover in Patients With Chronic Hepatitis C Responding to Treatment With rIFN- |
To test the possible synergistic antiviral efficacy of ribavirin in vivo, chronically HCV-infected patients were treated with 3 × 6 MU rIFN-![]() per week alone or in combination with 14 mg/kg ribavirin per day. The potential antiviral mechanism of ribavirin in HCV-infected patients is unknown. Assuming ribavirin inhibits virus release from infected cells, the half-life t1/2 = ln2/k1 in patients treated with rIFN-
per week alone or in combination with 14 mg/kg ribavirin per day. The potential antiviral mechanism of ribavirin in HCV-infected patients is unknown. Assuming ribavirin inhibits virus release from infected cells, the half-life t1/2 = ln2/k1 in patients treated with rIFN-![]() and ribavirin should become longer compared with patients treated with rIFN-
and ribavirin should become longer compared with patients treated with rIFN-![]() alone. However, the half-life of virus release from infected cells was clearly not prolonged in patients under combination therapy (t1/2 = ln2/k1= 1.99 ± 1.70) compared with patients treated with rIFN-
alone. However, the half-life of virus release from infected cells was clearly not prolonged in patients under combination therapy (t1/2 = ln2/k1= 1.99 ± 1.70) compared with patients treated with rIFN-![]() alone (t1/2 = ln2/k1 = 2.54 ± 2.10). In addition, half-lives of viral learance from serum were similar in patients treated with rIFN-
alone (t1/2 = ln2/k1 = 2.54 ± 2.10). In addition, half-lives of viral learance from serum were similar in patients treated with rIFN-![]() or rIFN-
or rIFN-![]() plus ribavirin (t1/2 = ln2/k2 = 0.23 ± 0.15 vs. 0.18 ± 0.19 days) (table 2).
plus ribavirin (t1/2 = ln2/k2 = 0.23 ± 0.15 vs. 0.18 ± 0.19 days) (table 2).
Hepatocyte damage and turnover can be estimated only by surrogate parameters such as transaminases that are released because of direct virus-related cytopathic and/or immune-mediated processes. In HCV-infected patients responding to antiviral therapy, the rapid decline of HCV RNA after initiation of treatment is accompanied by a slower decline of both serum ALT and serum AST toward normal levels. In patients responding to antiviral therapy, mean ALT activity before initiation of treatment (B0ALT) was 83.1 ± 56.1 U/L (3 × 3 MU rIFN-![]() /wk), 84.0 ± 49.7 U/L (3 × 6 MU rIFN-
/wk), 84.0 ± 49.7 U/L (3 × 6 MU rIFN-![]() /wk), and 58.7 ± 28.8 U/L (3 × 6 MU rIFN-
/wk), and 58.7 ± 28.8 U/L (3 × 6 MU rIFN-![]() /wk + 14 mg/kg ribavirin/day) (table 2). During antiviral therapy, ALT declined toward baseline levels (B
/wk + 14 mg/kg ribavirin/day) (table 2). During antiviral therapy, ALT declined toward baseline levels (B![]() ALT) of 8.9 ± 2.5 U/L, 14.6 ± 7.1 U/L, and 9.7 ± 3.4 U/L, respectively (Fig. 1). Results for ALT release from hepatocytes of patients treated with 3 × 3 MU rIFN-
ALT) of 8.9 ± 2.5 U/L, 14.6 ± 7.1 U/L, and 9.7 ± 3.4 U/L, respectively (Fig. 1). Results for ALT release from hepatocytes of patients treated with 3 × 3 MU rIFN-![]() /wk yield half-lives of t1/2ALT = ln2/k3 = 4.3 ± 4.4 days and for ALT degradation of t1/2ALT = ln2/k4 = 3.5 ± 1.5 days. In patients treated with 3 × 6 MU rIFN-
/wk yield half-lives of t1/2ALT = ln2/k3 = 4.3 ± 4.4 days and for ALT degradation of t1/2ALT = ln2/k4 = 3.5 ± 1.5 days. In patients treated with 3 × 6 MU rIFN-![]() /wk plus ribavirin, the half-life of ALT release was prolonged compared with patients treated with 3 × 6 MU rIFN-
/wk plus ribavirin, the half-life of ALT release was prolonged compared with patients treated with 3 × 6 MU rIFN-![]() /wk alone (t 1/2ALT= ln2/k3 = 7.8 ± 9.4 vs. 3.2 ± 2.5 days; P = .15), whereas half-lives of ALT degradation were similar in both groups (table 2). Kinetic data for AST revealed similar information (table 2).
/wk alone (t 1/2ALT= ln2/k3 = 7.8 ± 9.4 vs. 3.2 ± 2.5 days; P = .15), whereas half-lives of ALT degradation were similar in both groups (table 2). Kinetic data for AST revealed similar information (table 2).
DISCUSSION
HCV infection often progresses to chronic hepatitis, cirrhosis, and possibly hepatocellular carcinoma.25-27 IFN-![]() is the only approved treatment of chronic Hepatitis C; however, the sustained response rate with respect to detectable viremia is below 20%. Ribavirin is a non
is the only approved treatment of chronic Hepatitis C; however, the sustained response rate with respect to detectable viremia is below 20%. Ribavirin is a non![]() interferon-inducing nucleoside analogue with a broad spectrum of activity against RNA and DNA viruses, including those from the Flaviviridae family.4,8 Several studies have been conducted to evaluate ribavirin monotherapy in daily doses of 600 to 1,200 mg in the treatment of chronic Hepatitis C.3,4 Although the results consistently showed a decrease of transaminase levels, the biochemical response was neither associated with suppression of HCV RNA nor maintained after the drug was discontinued. Small pilot studies on combination of ribavirin and IFN-
interferon-inducing nucleoside analogue with a broad spectrum of activity against RNA and DNA viruses, including those from the Flaviviridae family.4,8 Several studies have been conducted to evaluate ribavirin monotherapy in daily doses of 600 to 1,200 mg in the treatment of chronic Hepatitis C.3,4 Although the results consistently showed a decrease of transaminase levels, the biochemical response was neither associated with suppression of HCV RNA nor maintained after the drug was discontinued. Small pilot studies on combination of ribavirin and IFN-![]() indicated a potential synergistic effect in the treatment of chronic Hepatitis C.5 This was confirmed in larger trials, particularly in terms of maintenance of long-term response.6,7 Kinetic analysis of viral turnover revealed a half-life of HCV in the order of a few hours. In the present study, minimum virus production and clearance per day was calculated to be approximately 1.8 × 1011 virions per day, confirming previously published data.14 As for human immunodeficiency virus-1,15,24,28 the vast majority of circulating HCV is supposed to derive from continuous rounds of de novo virus infection, replication and cell turnover, and not from cells chronically producing virions. The high turnover rates for both HCV and human immunodeficiency virus-1 explain the rapid generation of viral diversity and the opportunity for viral escape phenomena from the host immune surveillance.14,15,24,28
indicated a potential synergistic effect in the treatment of chronic Hepatitis C.5 This was confirmed in larger trials, particularly in terms of maintenance of long-term response.6,7 Kinetic analysis of viral turnover revealed a half-life of HCV in the order of a few hours. In the present study, minimum virus production and clearance per day was calculated to be approximately 1.8 × 1011 virions per day, confirming previously published data.14 As for human immunodeficiency virus-1,15,24,28 the vast majority of circulating HCV is supposed to derive from continuous rounds of de novo virus infection, replication and cell turnover, and not from cells chronically producing virions. The high turnover rates for both HCV and human immunodeficiency virus-1 explain the rapid generation of viral diversity and the opportunity for viral escape phenomena from the host immune surveillance.14,15,24,28
Mathematical analysis of viral decay data can be exploited to predict the mechanism of action and to compare the efficacy of antiviral compounds in vivo. Previous analysis of HCV RNA decay data were in favor of a predominant effect of rIFN-![]() on inhibition of de novo infection of susceptible cells and argued against a predominant antiviral effect of rIFN-
on inhibition of de novo infection of susceptible cells and argued against a predominant antiviral effect of rIFN-![]() on HCV release from infected cells. In the present study, we compared the antiviral efficacy of two different rIFN-
on HCV release from infected cells. In the present study, we compared the antiviral efficacy of two different rIFN-![]() dosages (3 × 3 MU and 3 × 6 MU per week). An initial decline of HCV RNA levels below the detection limit of the reverse-transcription polymerase chain reaction assay was observed in 10 of 26 (39%) and 10 of 19 patients (53%), respectively. Although the data suggest higher initial response rates in patients treated with 3 × 6 MU rIFN-
dosages (3 × 3 MU and 3 × 6 MU per week). An initial decline of HCV RNA levels below the detection limit of the reverse-transcription polymerase chain reaction assay was observed in 10 of 26 (39%) and 10 of 19 patients (53%), respectively. Although the data suggest higher initial response rates in patients treated with 3 × 6 MU rIFN-![]() per week, the difference was not significant (P = .52). In both treatment schedules, the most significant fit of the observed HCV-RNA decline after initiation of therapy was obtained assuming that the predominant antiviral effect of rIFN-
per week, the difference was not significant (P = .52). In both treatment schedules, the most significant fit of the observed HCV-RNA decline after initiation of therapy was obtained assuming that the predominant antiviral effect of rIFN-![]() is inhibition of de novo infection of susceptible cells (k
is inhibition of de novo infection of susceptible cells (k![]() 1 = 0). As in the previous study,14 the model assuming total inhibition of HCV replication by rIFN-
1 = 0). As in the previous study,14 the model assuming total inhibition of HCV replication by rIFN-![]() in infected cells (k1 = 0) showed an inappropriate fit of the observed data.
in infected cells (k1 = 0) showed an inappropriate fit of the observed data.
Assuming that the predominant effect of IFN is inhibition of de novo infection of susceptible cells, the HCV RNA concentration can only fall if the number of infected cells decays. Several lines of evidence support the hypothesis of a rapid hepatocyte turnover: 1) hepatocyte turnover can be estimated by surrogate parameters such as transaminases that are released because of direct virus-related cytopathic and/or immune-mediated processes. In HCV-infected patients responding to rIFN-![]() , the rapid decline of HCV RNA after initiation of treatment is accompanied by a similar decline of both serum ALT and serum AST toward normal levels.14 The calculated half-lives of virus release and transaminases are similar, indicating that both virus and hepatocytes have high turnover rates; 2) the liver of HCV-infected patients is infiltrated by a large number of cytotoxic T lymphocytes29,30; 3) in liver tissue of patients with chronic Hepatitis C, a high expression of Fas antigen as an inducer of apoptosis is observed31; and 4) regeneration of the liver, e.g., after partial resection is extremely rapid.32 A precedent for a high hepatocyte turnover rate exists in patients with chronic Hepatitis B. Nowak et al. calculated that approximately 109 hepatocytes are killed and replenished every day.16 Calculation of kinetic parameters revealed similar half-lifes of approximately 2.5 days for virus release from infected cells in patients treated with either 3 × 3 MU or 3 × 6 MU rIFN-
, the rapid decline of HCV RNA after initiation of treatment is accompanied by a similar decline of both serum ALT and serum AST toward normal levels.14 The calculated half-lives of virus release and transaminases are similar, indicating that both virus and hepatocytes have high turnover rates; 2) the liver of HCV-infected patients is infiltrated by a large number of cytotoxic T lymphocytes29,30; 3) in liver tissue of patients with chronic Hepatitis C, a high expression of Fas antigen as an inducer of apoptosis is observed31; and 4) regeneration of the liver, e.g., after partial resection is extremely rapid.32 A precedent for a high hepatocyte turnover rate exists in patients with chronic Hepatitis B. Nowak et al. calculated that approximately 109 hepatocytes are killed and replenished every day.16 Calculation of kinetic parameters revealed similar half-lifes of approximately 2.5 days for virus release from infected cells in patients treated with either 3 × 3 MU or 3 × 6 MU rIFN-![]() per week. In a previous study, the half-life for virus release in patients treated with 3 × 3 MU rIFN-
per week. In a previous study, the half-life for virus release in patients treated with 3 × 3 MU rIFN-![]() per week was t1/2 = 2.7 ± 1.3.14 The more pronounced decline of HCV RNA levels as observed in patients treated with 3 × 6 MU compared with 3 × 3 MU rIFN-
per week was t1/2 = 2.7 ± 1.3.14 The more pronounced decline of HCV RNA levels as observed in patients treated with 3 × 6 MU compared with 3 × 3 MU rIFN-![]() per week was caused by significantly faster clearance of HCV from serum. This suggests that IFN-
per week was caused by significantly faster clearance of HCV from serum. This suggests that IFN-![]() exerts a dose-dependent effect on (possibly) antigen-specific degradation of free virus in patients with chronic Hepatitis C (e.g., phagocytosis of HCV by macrophages, modulation of antibody production).33 Dose-dependent IFN-
exerts a dose-dependent effect on (possibly) antigen-specific degradation of free virus in patients with chronic Hepatitis C (e.g., phagocytosis of HCV by macrophages, modulation of antibody production).33 Dose-dependent IFN-![]() effects on HCV RNA clearance have previously also been quantified by Lam et al.34 As recently pointed out,35 the combined effects of pharmacological and intracellular delays, the clearance of free virus particles, and the decay of infected cells may cause the half-life of serum virus to be overestimated. In our study, time intervals between blood samples were too large to assess such effects.
effects on HCV RNA clearance have previously also been quantified by Lam et al.34 As recently pointed out,35 the combined effects of pharmacological and intracellular delays, the clearance of free virus particles, and the decay of infected cells may cause the half-life of serum virus to be overestimated. In our study, time intervals between blood samples were too large to assess such effects.
In addition, we investigated the in vivo antiviral efficacy of IFN-![]() in combination with ribavirin. The rate constants k1 and k2 of HCV turnover were similar in patients under combination therapy and patients treated with rIFN-
in combination with ribavirin. The rate constants k1 and k2 of HCV turnover were similar in patients under combination therapy and patients treated with rIFN-![]() in the absence of ribavirin. This apparently excludes a direct synergistic antiviral effect of rIFN-
in the absence of ribavirin. This apparently excludes a direct synergistic antiviral effect of rIFN-![]() and ribavirin. These kinetic data are well in accordance with clinical trials showing similar virological response rates at the end of treatment in patients treated with rIFN-
and ribavirin. These kinetic data are well in accordance with clinical trials showing similar virological response rates at the end of treatment in patients treated with rIFN-![]() with and without ribavirin.7 The improved sustained biochemical and virological response rates after discontinuation of combination therapy are indicative of immunmodulatory effects of ribavirin, as recently described in animal models.36
with and without ribavirin.7 The improved sustained biochemical and virological response rates after discontinuation of combination therapy are indicative of immunmodulatory effects of ribavirin, as recently described in animal models.36
In agreement with the clinical observation that serum transaminase levels decline during ribavirin treatment,3,4 we found the mean rate constant k3 for ALT and AST release from hepatocytes diminished in patients treated with IFN-![]() and ribavirin. However, the differences of k3 between both treatment groups did not reach statistical significance (P = .15 and P =.33 for k3ALT and k3AST, respectively). The overall observation under ribavirin therapy is an accelerated clearance of transaminases caused by a change in the production-elimination ratio.
and ribavirin. However, the differences of k3 between both treatment groups did not reach statistical significance (P = .15 and P =.33 for k3ALT and k3AST, respectively). The overall observation under ribavirin therapy is an accelerated clearance of transaminases caused by a change in the production-elimination ratio.
In conclusion, the data of the present study indicate that higher IFN-![]() doses enhance viral clearance from serum. Combination therapy with ribavirin, however, has no direct synergistic antiviral effect in the treatment of chronic Hepatitis C with 6 MU rIFN-
doses enhance viral clearance from serum. Combination therapy with ribavirin, however, has no direct synergistic antiviral effect in the treatment of chronic Hepatitis C with 6 MU rIFN-![]() three times per week. According to the reduction of transaminases as surrogate markers for hepatocyte damage, ribavirin is likely to have immunmodulatory and/or anti-inflammatory effects in patients with chronic Hepatitis C.
three times per week. According to the reduction of transaminases as surrogate markers for hepatocyte damage, ribavirin is likely to have immunmodulatory and/or anti-inflammatory effects in patients with chronic Hepatitis C.
Footnotes
Abbreviations: rIFN-![]() , recombinant interferon alfa; HCV, Hepatitis C virus; ALT, alanine transaminase; AST, aspartate transaminase.
, recombinant interferon alfa; HCV, Hepatitis C virus; ALT, alanine transaminase; AST, aspartate transaminase.
Supported by the Bundesministerium für Bildung, Wissenschaft, Forschung und Technologie (BMBF)
Received August 20, 1997; accepted April 24, 1998.
Address reprint requests to: Prof. Dr. med. Stefan Zeuzem, Medizinische Klinik II, Zentrum der Inneren Medizin, Klinikum der Johann Wolfgang Goethe-Universität, Theodor-Stern-Kai 7, D-60590 Frankfurt a.M., Germany. Fax: 49-69-6301-6448.
REFERENCES
- Poynard T, Leroy V, Cohard M, Thevenot T, Mathurin P, Opolon P, Zarski JP. Meta-analysis of interferon randomized trials in the treatment of viral Hepatitis C: effects of dose and duration. HEPATOLOGY 1996; 24: 778-789.
- Balart LA, Perrillo R, Roddenberry J, Regenstein F, Shim K-S, Shieh YSC, Taylor B, et al. Hepatitis C RNA in liver of chronic Hepatitis C patients before and after interferon alfa treatment. Gastroenterology 1993; 104: 1472-1477.
- Di Bisceglie AM, Conjeevaram HS, Fried MW, Sallie R, Park Y, Yurdaydin C, Swain M, et al. Ribavirin as therapy for chronic Hepatitis C. A randomized, double-blind, placebo-controlled trial. Ann Intern Med 1995; 123: 897-903.
- Dusheiko G, Main J, Thomas H, Reichard O, Lee C, Dhillon A, Rassam S, et al. Ribavirin treatment for patients with chronic Hepatitis C: results of a placebo-controlled study. J Hepatol 1996; 25: 591-598.
- Brillanti S, Garson J, Foli M, Whitby K, Deaville R, Masci C, Miglioli M, et al. A pilot study of combination therapy with ribavirin plus interferon alfa for interferon alfa-resistant chronic Hepatitis C. Gastroenterology 1994; 107: 812-817.
- Lai MY, Kao JH, Yang PM, Wang JT, Chen PJ, Chan KW, Chu JS, et al. Long-term efficacy of ribavirin plus interferon alfa in the treatment of chronic Hepatitis C. Gastroenterology 1996; 111: 1307-1312.
- Reichard O, Norkrans G, Frydén A, Braconier J-H, Sönnerborg A, Weiland O, for the Swedish Study Group. Randomised, double-blind, placebo-controlled trial of interferon
 -2b with and without ribavirin for chronic Hepatitis C. Lancet 1998; 351: 83-87.
-2b with and without ribavirin for chronic Hepatitis C. Lancet 1998; 351: 83-87. - Sidwell R, Huffman JH, Khare L, Allen LB, Witlowski JT, Robins RK. Broad-spectrum activity of virazole: 1-beta-D-ribofuranosyl-1,2,4-triazole-3-carboxamide. Science 1972; 117: 705-706.
- Rüster B, Zeuzem S, Roth WK. Quantification of Hepatitis C virus RNA by competitive reverse transcription and polymerase chain reaction using a modified Hepatitis C virus RNA transcript. Anal Biochem 1995; 224: 597-600.
- Roth WK, Lee J-H, Rüster B, Zeuzem S. Comparison of two quantitative Hepatitis C virus reverse transcriptase PCR assays. J Clin Microbiol 1996; 34: 261-264.
- Zeuzem S, Franke A, Lee J-H, Herrmann G, Rüster B, Roth WK. Phylogenetic analysis of Hepatitis C virus isolates and their correlation to viremia, liver function tests, and histology. HEPATOLOGY 1996; 24: 1003-1009.
- Simmonds P, Holmes EC, Cha TA, Chan SW, McOmish F, Irvine B, Beall E, et al. Classification of Hepatitis C virus into six major genotypes and a series of subtypes by phylogenetic analysis of the NS-5 region. J Gen Virol 1993; 74: 2391-2399.
- Lee J-H, Roth WK, Zeuzem S. Evaluation and comparison of different Hepatitis C virus genotyping and serotyping assays. J Hepatol 1997; 26: 1001-1009.
- Zeuzem S, Schmidt JM, Lee J-H, Rüster B, Roth WK. Effect of interferon alfa on the dynamics of Hepatitis C virus turnover in vivo. HEPATOLOGY 1996; 23: 366-371.
- Wei X, Ghosh SK, Taylor ME, Johnson VA, Emini EA, Deutsch P, Lifson JD, et al. Viral dynamics in human immunodeficiency virus type 1 infection. Nature 1995; 373: 117-122.
- Nowak MA, Bonhoeffer S, Hill AM, Boehme R, Thomas HC, McDade H. Viral dynamics in Hepatitis B virus infection. Proc Natl Acad Sci U S A 1996; 93: 4398-4402.
- Ballardini G, Groff P, Giostra F, Francesconi R, Miniero R, Ghetti S, Zauli D, et al. Hepatocellular codistribution of c100, c33, c22, and NS5 Hepatitis C virus antigens detected by using immunopurified polyclonal spontaneous human antibodies. HEPATOLOGY 1995; 21: 730-734.
- Lau GK, Davis GL, Wu SP, Gish RG, Balart LA, Lau JY. Hepatic expression of Hepatitis C virus RNA in chronic Hepatitis C: a study by in situ reverse-transcription polymerase chain reaction. HEPATOLOGY 1996; 23: 1318-1323.
- Nelder JA, Mead R. A simplex method for function minimization. Computer Journal 1965; 7: 308-313.
- Johnson ML. Evaluation and propagation of confidence intervals in nonlinear, asymmetrical variance spaces: analysis of ligand-binding data. Biophys J 1983; 44: 101-106.
- Waits DG. Parameter estimates from nonlinear models. Methods Enzymol 1994; 240: 23-36.
- Sokal RR, Rohlf FJ. Biometry–Statistical Tables, 2nd New York: W.H. Freeman , 1981.
- Press WH, Flannery BP, Teukolsky SA, Vetterling WT. Numerical Recipes. Cambridge: Cambridge University Press , 1989.
- Ho DD, Neumann AU, Perelson AS, Chen W, Leonard JM, Markowitz M. Rapid turnover of plasma virions and CD4 lymphocytes in HIV-1 infection. Nature 1995; 373: 123-126.
- lter MJ, Margolis HS, Krawczynski K, Judson FN, Mares A, Alexander WJ, Hu PY, et al. The natural history of community-acquired Hepatitis C in the United States. N Engl J Med 1992; 327: 1899-1905.
- Tong MJ, El-Farra NS, Reikes AR, Co RL. Clinical outcomes after transfusion-associated Hepatitis C. N Engl J Med 1995; 332: 1463-1466.
- aito I, Miyamura T, Ohbayashi A, Harada H, Katayama T, Kikuchi S, Watanabe Y, et al. Hepatitis C virus infection is associated with the development of hepatocellular carcinoma. Proc Natl Acad Sci U S A 1990; 87: 6547-6549.
- Perelson AS, Neumann AU, Markowitz M, Leonard JM, Ho DD. HIV-1 dynamics in vivo: virion clearance rate, infected cell life-span, and viral generation time. Science 1996; 271: 1582-1586.
- Koziel MJ, Dudley D, Wong JT, Dienstag J, Houghton M, Ralston R, Walker BD. Intrahepatic cytotoxic T lymphocytes specific for Hepatitis C virus in persons with chronic hepatitis. J Immunol 1992; 149: 3339-3344.
- Ballardini G, Groff P, Pontisso P, Giostra F, Francesconi R, Lenzi M, Zauli D, et al. Hepatitis C virus (HCV) genotype, tissue HCV antigens, hepatocellular expression of HLA-A,B,C, and intercellular adhesion-1 molecules. J Clin Invest 1995; 95: 2067-2075.
- Hiramatsu N, Hayashi N, Katayama K, Mochizuki K, Kawanishi Y, Kasahara A, Fusamoto H, et al. Immunohistochemical detection of fas antigen in liver tissue of patients with chronic Hepatitis C. HEPATOLOGY 1994; 19: 1354-1359.
- Fausto N, Webber EM. In: Arias IM, Boyer JL, Fausto N, Jakoby WB, Schachter D, Shafritz DA, eds. The Liver: Biology and Pathobiology, 3rd ed. New York: Raven , 1994.
- Gutterman JU. Cytokine therapeutics: lessons from interferon
 . Proc Natl Acad Sci U S A 1994; 91: 1198-1205
. Proc Natl Acad Sci U S A 1994; 91: 1198-1205 - Lam NP, Neumann AU, Gretch DR, Wiley TE, Perelson AS, Layden AJ. Dose-dependent acute clearance of Hepatitis C genotype 1 virus with interferon alfa. HEPATOLOGY 1997; 26: 226-231.
- Herz AVM, Bonhoeffer S, Anderson RM, May RM, Nowak MA. Viral dynamics in vivo: limitations on estimates of intracellular delay and virus decay. Proc Natl Acad Sci U S A 1996; 93: 7247-7251.
- Ning Q, Brown D, Parodo J, Cattral M, Fung L, Liu M, Rotstein O, Levy G. Ribavirin inhibits viral induced macrophage production of tumor necrosis factor, interleukin 1 and procoagulant activity and preserves Th1 cytokine production, but inhibits Th2 cytokine response [Abstract]. HEPATOLOGY 1996; 24: 355A.
Copyright © 1998 by the American Association for the Study of Liver Diseases.